Protein biosynthesis steps, site, importance, inhibitors and Protein maturation
Protein biosynthesis is a core biological process, It can be divided broadly into two phases – transcription and translation, It takes place inside cells to balance the loss of cellular proteins (via degradation or export) through the production of new proteins, Proteins can perform critical functions as enzymes, structural proteins or hormones and therefore, they are crucial biological components.
Protein biosynthesis (Synthesis)
Protein biosynthesis (protein synthesis) can be described in 3 phases: initiation, elongation, and termination, the protein sequence is synthesized and read from the amino terminus to the carboxy terminus.
Initiation
For initiation of protein biosynthesis, there must be:
- mRNA
- rRNA
- tRNA
- Eukaryotic initiation factors (eIFs).
- GTP, ATP, and different amino acids.
tRNA charging
tRNA charging means the recognition & attachment of the specific amino acid to the 3´ hydroxyl adenosine terminus (to the sugar) of tRNA in an ester linkage.
Initiation involves 4 steps:
- Ribosomal dissociation: the 80 S eukaryotic ribosome is dissociated into 40 S and 60 S subunits, eIF-3 and eIF-1 bind to 40 S subunits thus preventing reassociation between the 2 subunits.
- Formation of 43 S preinitiation complex: The first step in this process involves the binding of GTP by eIF-2, this binary complex then binds to Met-tRNA ( a tRNA specifically involved in binding to the initiation codon AUG), this ternary complex binds to the 40 S ribosomal subunit to form preinitiation complex.
- Formation of 48 S initiation complex: mRNA binds to the preinitiation complex to form the 48 S initiation complex with the hydrolysis of ATP to ADP + Pi, this initiation complex scans the mRNA for a suitable initiation codon.
- The formation of 80 S initiation complex: The binding of 60 S ribosomal subunit to the 48 S initiation complex involves the hydrolysis of the GTP bound to eIF-2, this reaction results in the release of the initiation factors bound to 48 S initiation complex (these factors are recycled) and rapid association of 40 S and 60 S subunits to form the 80 S ribosome, at this point, Met-tRNA is on the peptidyl (P) site of the ribosome, ready for the elongation cycle to commence.
Elongation
It’s a cyclic process involving 3 steps:
- Binding of aminoacyl-tRNA to the A site: In the complete 80 S ribosome formed during initiation, the aminoacyl (A) site is free, the binding of the proper aminoacyl tRNA in the A site requires proper codon recognition and activation of aminoacyl tRNA by binding of eukaryote factor-1 (e EF-1) and GTP (ternary complex). when aminoacyl tRNA binds to A site, GTP is hydrolyzed and e EF-1 is released.
- Peptide bond formation: the α-amino group of the aminoacyl tRNA in the A site attacks the carboxyl group of the peptidyl-tRNA occupying the P site, this reaction is catalyzed by peptidyl transferase of the 60 S subunit (this is an example of ribozyme activity), the reaction results in attachment of the growing peptide chain to the tRNA in the A site.
- Translocation: upon the removal of the peptidyl moiety from the tRNA in the P site, the discharged tRNA quickly dissociates from the P site, eEF-2 and GTP are responsible for the translocation of the newly formed peptidyl tRNA at the A site into the empty P site. The translocation of the newly formed peptidyl tRNA, and its corresponding codon into the P site then frees the A site for another cycle of amino acyl-tRNA codon recognition and elongation.
The formation of one peptide bond requires the energy resulting from hydrolysis of 4 high energy phosphate bonds:
- Charging of tRNA with aminoacyl moiety requires the hydrolysis of an ATP to an AMP.
- The entry of amino tRNA into the A site requires one GTP hydrolysis to GDP.
- The translocation of the newly formed peptidyl-tRNA in the A site into the P site results in the hydrolysis of one GTP to GDP.
Termination
After many cycles of elongation, the non-sense or termination code of mRNA (UAA, UAG, UGA) appears in the A site. Normally, there is no tRNA with an anticodon capable of recognizing such termination signal, releasing factors (eRFs) can recognize the termination signals in the A site.
eRF, GTP, and peptidyl transferase promote the hydrolysis of the bond between the peptide and the tRNA occupying the P site, then the 80 S ribosome dissociates into 40 S and 60 S subunits and mRNA, tRNA, eRF, GDP & Pi are released.
Site of protein biosynthesis
Protein biosynthesis takes place on ribosomes, many ribosomes can assemble to translate a single mRNA molecule forming a polysome. Polysomes are either found free in the cytoplasm or attached to the rough endoplasmic reticulum, depending on the protein being translated, the free polysomes are responsible for the synthesis of proteins required for intracellular functions, while attached polysomes synthesize integral membrane proteins and proteins to be exported.
Protein maturation
Newly synthesized polypeptide chains undergo folding and posttranslational processing. These modifications may result in their activation to functional form, localization in a subcellular compartment, or their secretion from the cell.
I- Protein folding
As proteins emerge from ribosomes, they fold into three-dimensional conformations that are essential for their subsequent biologic activity. Molecular chaperones are proteins that assist in the process of proper protein folding, misfolded proteins are targeted for destruction.
II- Post-translational processing
(1) Proteolysis: proteolysis of proteins is a common maturation step, amino-terminal, carboxy-terminal or internal sequences can be removed from the protein.
Examples:
- Removal of amino-terminal Met residues of most eukaryotic proteins.
- Removal of signal peptides by signal peptidases.
(It’s a signal sequence which directs the protein to its appropriate location in the cell is often removed by signal peptidase after the protein has reached its final site).
(2) Modifications of individual amino acids:
Examples:
- Hydroxylation of proline and lysine to form hydroxyproline and hydroxylysine is important for collagen synthesis.
- Phosphorylation of serine, threonine, or tyrosine by ATP-requiring protein kinases regulates the activity of many enzymes e.g. glycogen phosphorylase.
- γ-Carboxylation of glutamic acid residues in prothrombin to form γ-carboxy glutamate; allows the binding of calcium ions and leads to the formation of the blood clots.
(3) Addition of certain groups:
Attachment of carbohydrate side chains (Glycosylation) to proteins forms glycoproteins, in some glycoproteins, the carbohydrate side chain is attached to asparagine residues (N-linked), in others to serine and threonine residues (O-linked).
Inhibitors of protein biosynthesis
- Effective antibiotics must interact specifically with prokaryotic ribosomes to inhibit bacterial protein synthesis without interaction with components of eukaryotic ribosomes (e.g. streptomycin, tetracycline, chloramphenicol, and erythromycin).
– Streptomycin inhibits the initiation of prokaryotic protein biosynthesis.
– Tetracyclines block the A site on the prokaryotic ribosome, thus inhibit the binding of aminoacyl tRNA.
– Chloramphenicol blocks peptidyl transfer on ribosomes.
– Erythromycin can inhibit the translocation reaction of prokaryotic protein synthesis. - Other antibiotics are not clinically useful because they can inhibit protein synthesis in both prokaryotes as eukaryotes (as puromycin) or only in eukaryotes (as cycloheximide).
- Some toxins inhibit eukaryotic protein synthesis (as diphtheria toxins and ricin).
DNA Repair types, definition & importance, Protein Biosynthesis & Genetic Code
DNA replication steps & rules, DNA polymerase enzymes & RNA primer synthesis
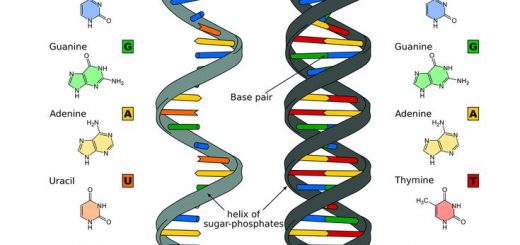
Structure of nucleic acids, Deoxyribonucleic acid (DNA) & Ribonucleic acid (RNA)
Importance of Nucleosides, Nucleotides, Purines, Pyrimidines & Sugars of nucleic acids
Gene mutations types, causes, examples & Regulation of Gene Expression