DNA Repair types, definition and importance, Protein Biosynthesis and Genetic Code
DNA repair is defined as the cellular responses which are associated with the restoration of the normal base-pair sequence and structure of damaged DNA. DNA repair depends on the cell type, the age of the cell, and the extracellular environment, Normal metabolic activities & environmental factors such as radiation cause DNA damage and can eliminate the cell’s ability to transcribe the gene that the affected DNA encodes.
DNA repair
DNA repair allows the cell to identify & correct damage to the DNA molecules that encode its genome. The best way to illustrate the importance of DNA repair is to consider the effects of unrepaired DNA damage. The most serious outcome is a change in the base sequence of DNA. which, if replicated and transmitted to future cell generations, becomes permanent. A permanent change in the nucleotide sequence of DNA is called a mutation. In mammals, there is a strong correlation between the accumulation of mutations and cancer.
Types of DNA repair
- Mismatch repair.
- Base excision repair.
- Nucleotide excision repair.
- Double-strand break repair.
Base excision repair of DNA; The enzyme uracil DNA glycosylase removes the uracil created by the spontaneous deamination of cytosine in the DNA. An endonuclease cuts the backbone near the defect, then, after an endonuclease removes a few bases, the defect is filled in by the action of a repair polymerase and the strand is rejoined by a ligase.
- Mismatched base (G2) is due to DNA replication errors, and repair enzymes are DNA polymerase & DNA ligase.
- Cytosine deamination to uracil (G1) is due to spontaneous/heat/chemicals, and repair enzymes are DNA polymerase & DNA ligase.
- Thymine dimers (G1) is due to UV radiation, and repair enzymes are DNA polymerase & DNA ligase.
- Double-strand break (G1) is due to Ionizing radiation and oxidation free radicals, and the repair enzyme is DNA ligase.
Xeroderma pigmentosum is an autosomal recessive disorder, characterized by extreme sensitivity to sunlight, ulceration, and skin cancer. The most common deficiency occurs in the excision endonuclease enzyme.
Gene expression
Synthesis of protein under the influence of genes is called gene expression. It includes RNA transcription and protein synthesis (translation).
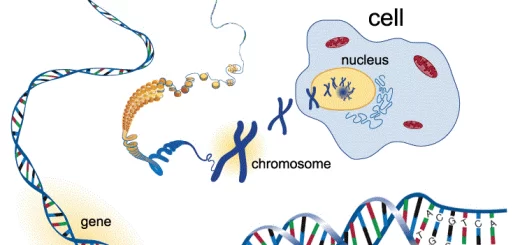
- Gene: is a DNA segment that contains well-defined genetic information and codes for an individual function e.g. polypeptide chain.
- Genome: means total DNA content of a cell (=total number of genes in one cell).
Transcription
Transcription is the process of synthesis of RNA from a DNA template by RNA polymerase enzymes and a number of associated proteins. For any particular gene, only one strand of the DNA molecule, called the template strand, is copied by RNA polymerase. The other strand is referring as coding (antitemplate strand) of that gene.
The coding strand is not used during transcription. It corresponds exactly to the sequence of the primary RNA transcript, except that RNA contains uracil instead of the thymine found in the DNA coding strand. DNA-dependent RNA polymerase elongates an RNA strand by adding ribonucleotide units to the strand 3´-hydroxyl end and builds RNA in the 5´→3´ direction.
Steps of RNA synthesis
The general steps required to synthesize the primary transcript (i.e a newly synthesized RNA) are initiation, elongation, and termination.
Initiation
Initiation protein factors and RNA polymerase are needed for the initiation process. RNA polymerase recognizes and binds to the promoter region (The promoter is the binding site for RNA polymerase on the double-stranded DNA. They have conserved or consensus sequences). The promoter identifies the start site for transcription and orients the enzyme on the template strand.
RNA polymerase then separates the two strands of DNA as it reads the base sequence of the template strand. Transcription begins at the +1 base pair. (initiation protein factors are released as soon as transcription is initiated).
The first nucleotide (always purine) is associated with the initiation site. Except for the first nucleoside triphosphate, subsequent nucleotides are added to the 3´-hydroxyl of the preceding nucleotide forming a phosphodiester bond and pyrophosphates are released.
Elongation
RNA polymerase continues moving along the template strand in the 3´→5´ direction, synthesizing RNA in 5´→3´ direction (elongation protein factors are needed in eukaryotes).
Termination
RNA polymerase eventually reaches a transcription termination signal, at that point, it will stop transcription and release the completed RNA molecule (primary transcript).
Post-transcriptional modification of RNA (RNA processing)
The immediate product of transcription is a precursor RNA molecule, the primary transcript, which is modified subsequently to a mature functional molecule. The primary transcript is copied from a linear segment of DNA, a transcription unit, between specific initiation and termination sites.
Processing of eukaryotic mRNA precursor (heterogeneous nuclear RNA; hn RNA):
Capping
The cap (7-methyl guanosine) is added to 5´ end while the RNA molecule is still being synthesized. 7-methyl guanosine is joined to the 5´ end of mRNA in an unusual 5´, 5´- triphosphate linkage. All mRNA is capped to be functional in protein biosynthesis.
Significance of the cap structure: serves as a ribosome binding site (for translation initiation). It protects the 5´ end of mRNA from attack by 5´→3´ exonuclease (for mRNA stability.
Polyadenylationnucleus
Poly (A) polymerase enzyme adds poly (A) tail to the 3´ end of mRNA (which is short at first about 20A residues, then subsequently extended to as much as 200 A residues).
Significance of the poly (A) tail: protects the 3´ end of mRNA from 3´→5 exonuclease attack (for mRNA stability). It aids in its transport from the nucleus to the cytoplasm.
Splicing
The coding portions, exons (the sequence of a gene that is represented as mRNA) of most eukaryotic genes are interrupted by intervening sequences; introns (the sequence of a gene that is transcribed but excised before translation).
Splicing means the removal of introns from the primary transcript in the nucleus and then ligation of exons to form mature functional m RNA. The mature mRNA is then transported to the cytoplasm where it is translated into protein.
Processing of eukaryotic transfer RNA precursor (pre-tRNA)
Eukaryotic tRNA genes are all transcribed by RNA polymerase III. The primary transcript (pre-tRNA) requires post-transcriptional processing to be mature functional rRNA such as:
- Folding and base pairing to generate its characteristic shape.
- Removal of excess nucleotides from the 5´ and 3´ ends (cleavage).
- Removal of introns (splicing).
- Addition of the CCA sequence at 3´ end.
- Modification of some bases by methylation, deamination, or reduction.
Processing of eukaryotic ribosomal RNA precursor (pre- rRNA)
The ribosomal subunits assemble in the nucleolus as rRNA pieces combine with ribosomal proteins. Eukaryotic ribosomal subunits are 60 S and 40 S. They join during protein synthesis to form the whole 80 S ribosome.
Protein Biosynthesis
The flow of genetic information from DNA to protein: The genetic information of the cell is stored and transmitted in the nucleotide sequences of DNA. Expression of this genetic information involves two stages:
- The first stage is the transcription to form mRNA that carries specific and precise messages in the form of codons from DNA to the cytoplasmic sites of protein synthesis.
- The second stage is the translation of the nucleotide sequence of an mRNA (Codons) into an amino acid sequence of a protein.
Each codon consists of a sequence of nucleotides i.e. it is a triplet code. The collection of these codons makes up the genetic code. Thus, protein biosynthesis is also called translation because it involves the translation of information from the 4-letter language and structure of nucleic acid into the 20-letter language and structure of proteins.
Components of the translational process
- mRNA as a carrier of genetic information.
- rRNA as an adapter molecule, which recognizes an amino acid on one end and its corresponding codon on the other end.
- Ribosomes as the molecular machines coordinating the interaction between mRNA, tRNA, the enzymes, and the protein factors required for protein synthesis.
Genetic Code
It is the relation between the sequence of nucleotides in DNA (or in m RNA) and the sequence of amino acids in a polypeptide chain. The terms first, second, and third nucleotide refer to the individual nucleotides of a triplet codon. U, uridine nucleotide, C, cytosine nucleotide; A, adenine nucleotide; G, guanine nucleotide; Met, chain initiator codon; Term, chain terminator codon. AUG, which codes for Met, serves as the initiator codon in mammalian cells.
mRNA contains 4 different nucleotides (A, G, C, and U). Each amino acid is specified by one or more codons (nucleotide triplets). Thus, there are 4³ codons (i.e. 64 codewords). Three codons (UAA, UAG, UGA) do not specify any amino acid, and function as the stop (termination or non-sense) codons in protein synthesis. The remaining 61 codons code for the 20 amino acids used in protein synthesis. There is one start codon (initiation codon), AUG, coding for methionine (Met). The genetic information along mRNA is read from 5´ to 3´ direction.
Characters of genetic code
Genetic code is degenerate i.e. multiple codons can code for the same amino acid except tryptophan and methionine. The 3rd nucleotide in a codon is less important than the other two in determining the specific amino acid to be incorporated (Wobble theory). Wobble, therefore allows a single tRNA to read (translate) several different codons for the same amino acid.
For example, the two codons for arginine (AGA and AGG) can bind to the same anticodon (UCU). (Note that the direction of reading the anticodon of tRNA is from 3´ to 5´, while reading the genetic code is from 5´ to 3´ direction).
Genetic code is unambiguous i.e. each codon specifies no more than one amino acid. Genetic code is non-overlapping and Commaless i.e. the reading of the genetic code does not involve any overlap of codons, and the message is read in a continuing sequence of nucleotide triplets until a termination sequence is reached. Genetic code is universal i.e. the same code words are used in all organisms (pro-and eukaryotes) with the exception in mitochondria.
Protein, Protein biosynthesis steps, site, importance, inhibitors & Protein maturation
DNA replication steps & rules, DNA polymerase enzymes & RNA primer synthesis
Structure of nucleic acids, Deoxyribonucleic acid (DNA) & Ribonucleic acid (RNA)
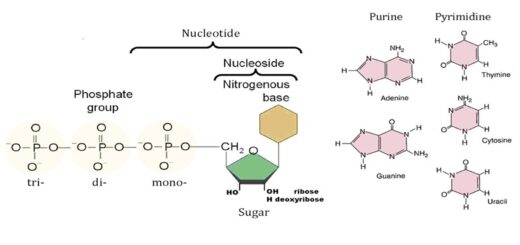
Importance of Nucleosides, Nucleotides, Purines, Pyrimidines & Sugars of nucleic acids
Regulation of the cell cycle, DNA synthesis phase, Interphase & Mitosis