DNA replication steps and rules, DNA polymerase enzymes and RNA primer synthesis
DNA replication is the process of DNA synthesis using parent DNA strands as a template. It aims at the formation of a copy of the parent DNA molecule for the daughter cell. DNA replication begins at specific locations of replication in the cell, and it produces two identical replicas of DNA from one original DNA molecule.
DNA replication
DNA replication is a biological process that occurs in all living organisms acting as the most essential part of biological inheritance. It is the process by which DNA makes a copy of itself during cell division.
Rules of DNA replication in eukaryotes
- DNA replication is semiconservative: Each DNA strand serves as a template for the synthesis of a new strand producing two DNA molecules, each with one new strand and one old strand.
- Replication begins at multiple origins and usually proceeds bidirectionally. Having multiple origins of replication provides a mechanism for rapidly replicating the great length of eukaryotic DNA molecules.
- Replication exhibits polarity: DNA synthesis proceeds in a 5´ → 3´ direction and is semi-discontinuous.
- Replication is very accurate: replication proceeds with an extraordinary degree of fidelity.
- DNA replication occurs in the nucleus during the synthetic (S) phase of the eukaryotic cell cycle. This phase is preceded and followed by two periods during which DNA is not synthesized (gap periods G1, and G2). During cell division (mitotic; M phase), each daughter cell receives one of the two identical DNA molecules.
During polymerization, discrimination between correct and incorrect nucleotides depends on base-pairing interactions (A=T and G≡C). Proofreading 3´→ 5´ exonuclease activity double-checks each nucleotide after it is added and removes the mispaired one. The additional accuracy is accounted for by a separate enzyme system that repairs the mismatched base pairs remaining after replication.
Requirements of DNA replication in eukaryotes
DNA is synthesized by DNA polymerases
DNA polymerases require the presence of a primer (i.e. oligonucleotide of RNA with free 3´ hydroxyl group), a template (i.e single-stranded DNA), and deoxyribonucleotides (d ATP, d CTP, d GTP, and d TTP) in order to function. The primer provides a site for the polymerization to begin. The free 3´ hydroxyl group of the primer acts as an acceptor for the first deoxyribonucleotide in the newly formed DNA strand.
DNA polymerases utilize one deoxyribonucleoside triphosphate as a source of the deoxyribonucleoside monophosphate for the growing DNA strand by the removal of pyrophosphate. The selection of the incoming deoxyribonucleotide is dependent upon proper base pairing with the template. The RNA-primed synthesis of DNA demonstrating the template function of the complementary strand of parental DNA.
In eukaryotes, there are at least 5 DNA polymerase enzymes (α, β, γ, δ and ε )
- DNA polymerase α has a Primase activity (for the synthesis of RNA primer); Polymerization activity (formation of phosphodiester bond), and No proofreading 3´ → 5´ exonuclease activity.
- DNA polymerase β functions in DNA repair (it has 5´→ 3´ exonuclease activity).
- DNA polymerase γ synthesizes mitochondrial DNA.
- DNA polymerase δ has Polymerization activity and Proofreading 3´→5´ exonuclease activity.
- DNA polymerase ε removes the primers of Okazaki fragments on the lagging strand. It has DNA repair and proofreading activities.
DNA is degraded by nucleases or DNases
They may be either exonucleases or endonucleases. Exonucleases degrade nucleic acids from one end of the molecule operating either from 5´→ 3´ or from 3´ →5´ direction of one strand of the double-stranded nucleic acids.
Endonucleases can begin to degrade at any internal site in a nucleic acid strand, reducing it to smaller and smaller fragments. Other enzymes (e.g. helicase, topoisomerase, and DNA ligase) and protein factors (e.g. origin binding proteins and single-stranded binding proteins) are required for the replication process.
Steps of DNA replication in eukaryotes
The synthesis of a DNA molecule can be divided into three stages: initiation, elongation, and termination.
Initiation
Identification of the origins of replication: Origins of replication in eukaryotes (e.g. yeast) are called replicators. These origins are located adjacent to an A-T-rich sequence that is easy to unwind. Origin recognition complex (ORC) is a set of proteins that can bind to these replicators.
The unwinding of double-stranded DNA: The interaction of these proteins with the replicators leads to the unwinding of DNA forming “replication bubbles”.
Formation of replication forks:
- When the two DNA strands unwind and separate, they form two replication forks at each origin. Replication of double-stranded DNA is bidirectional i.e. replication forks move in both directions away from the origin.
- DNA helicase binds to the unwound region near the replication fork and then moves into the neighboring double-stranded region, forcing the strands apart. Helicase uses energy from ATP to break the hydrogen bonds holding the base pairs together.
- As helicase unwinds the DNA at the replication forks, the DNA ahead of it becomes overwound and positive supercoils forms. Topoisomerase II removes these supercoils (by nicking both strands of DNA, passing the DNA strands through the nick, and then resealing both strands again).
- Eukaryotic single-stranded DNA binding protein binds to the single-stranded portion of each DNA strand, preventing the strands from reassociation and protecting them from degradation by nucleases.
RNA primer synthesis: DNA polymerase α with primase activity is responsible for the synthesis of a short RNA primer in 5´→3´ direction, beginning at the origin of each parental strand. The parental strand is used as a template for this process.
Elongation
The elongation phase of replication includes 2 distinct but related operations: leading and lagging strand synthesis.
Leading strand synthesis begins with the synthesis of RNA primer by DNA polymerase α at the replication origin. Deoxyribonucleotides are added to this primer by DNA polymerase δ. Leading strand synthesis then proceeds continuously, in the direction of replication fork movement.
Lagging strand synthesis is synthesized discontinuously in the opposite direction from the replication fork movement as a series of small fragments known as Okazaki fragments. Each Okazaki fragment is initiated by the synthesis of an RNA primer by polymerase α and then completed by the synthesis of DNA by DNA polymerase δ until reaching the next RNA primer.
Both leading and Okazaki fragments of lagging strands are synthesized from 5´→ 3´ direction. There is a leading and a lagging strand for each of the two replication forks. Then, DNA polymerase ε begins to remove the RNA primers. The gap is then filled by a polymerase (δ/ε).
Both DNA polymerase δ and ε can proofread their work using a 3´→5´ exonuclease activity. If DNA polymerase makes a mistake during DNA synthesis, the resulting unpaired base at the 3´ end of the growing strand is removed before synthesis continues.
Ligation of the newly synthesized DNA segments: DNA ligase seals the nicks between Okazaki fragments, converting them to a continuous strand of DNA. DNA ligase requires ATP to form a phosphodiester bond between the two nucleotides.
Termination
The termination of replication on linear eukaryotic chromosomes involves the synthesis of telomeres at the ends of the chromosome.
Telomeres
They are repetitive sequences TTAGGGs, these telomeres (Hexamers) cap the end of human chromosomes and are believed to prevent chromosomes from undergoing degradation. Telomeres are believed to contribute to the preservation of the normal aging process. With each round of replication in most normal cells, the telomeres are shortened.
During normal cell division, the length of telomeres is shortened, thus limiting the life span of the cell, Therefore, normal somatic cells after a certain number of cell division die due to the shortening of telomeres. So telomeres determine the proliferation capacity of human somatic cells.
Telomerase
It is an enzyme in eukaryotes used to maintain the telomeres. It contains a short RNA template complementary to the DNA telomere sequence, as well as telomerase reverse transcriptase activity. Telomerase is thus able to replace telomere sequences that would otherwise be lost during replication. Normally, telomerase activity is present only in embryonic cells, germ (reproductive) cells, and stem cells but not in somatic cells.
Cancer cells often have relatively high levels of telomerase, preventing the telomeres from becoming shortened and contributing to the immortality of malignant cells. After replication is complete, the parent and daughter strands reform double-stranded DNA.
In eukaryotic cells, the double-stranded DNA must precisely reform the chromatin structure including nucleosomes that existed prior to the onset of replication. The 2 identical sister chromatids are separated from each other when the cell divides during mitosis.
Quinolones are a family of drugs that block the action of prokaryotic topoisomerase thus preventing DNA replication and transcription. These drugs such as Nalidixic acid and ciprofloxacin act as antibiotics. While inhibitors of eukaryotic topoisomerases as Etoposide are becoming useful as anticancer agents.
DNA Repair types, definition & importance, Protein Biosynthesis & Genetic Code
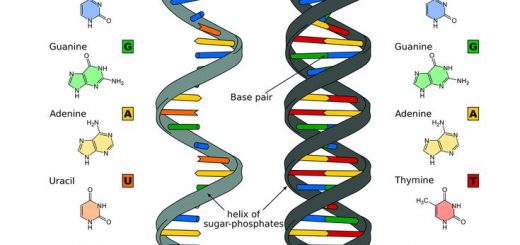
Structure of nucleic acids, Deoxyribonucleic acid (DNA) & Ribonucleic acid (RNA)
Importance of Nucleosides, Nucleotides, Purines, Pyrimidines & Sugars of nucleic acids
Regulation of the cell cycle, DNA synthesis phase, Interphase & Mitosis
Structure of Nucleic Acid (DNA), Enzymes, DNA replication and DNA repair mechanism
Cell division types, Mitosis, Meiosis, Reductional division and Equational division